This page was produced as an assignment for Genetics 677, an undergraduate course at UW-Madison.
Protein Phylogeny
Analysis
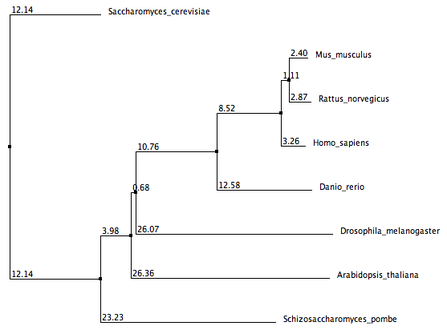
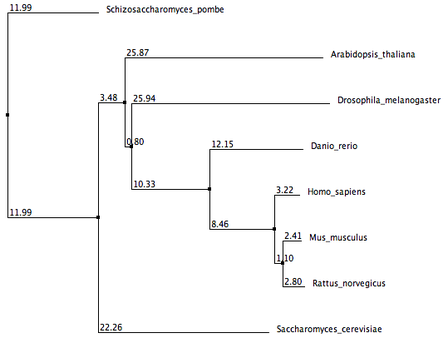
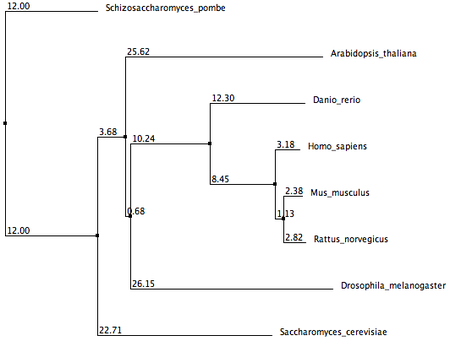
These phylogenetic trees were generated using three seperate online alignment tools, each with a different algorithm. The three tools used are ClustalW2, MUSCLE, and T-Coffee. Each algorithm came up with different lengths and orders of branches, but there are still some important similarities to note. In all phylogenetic trees, Mus musculus (Mouse) and Rattus norvegicus (Rat) share the closest common node to the human MSH2. This supports previous gene and protein alignment data that suggest mouse and rat animal models would be most appropriate for studying the human MSH2 protein. Schizosaccharomyces pombe homolog also seems to be the least related homolog to the human MSH2 protein in all the phylogenetic trees.
These phylogenetic trees were generated using three seperate online alignment tools, each with a different algorithm. The three tools used are ClustalW2, MUSCLE, and T-Coffee. Each algorithm came up with different lengths and orders of branches, but there are still some important similarities to note. In all phylogenetic trees, Mus musculus (Mouse) and Rattus norvegicus (Rat) share the closest common node to the human MSH2. This supports previous gene and protein alignment data that suggest mouse and rat animal models would be most appropriate for studying the human MSH2 protein. Schizosaccharomyces pombe homolog also seems to be the least related homolog to the human MSH2 protein in all the phylogenetic trees.