This page was produced as an assignment for Genetics 677, an undergraduate course at UW-Madison.
Protein Domains
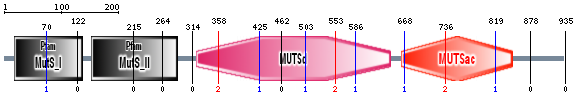
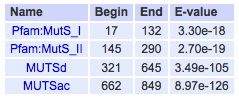
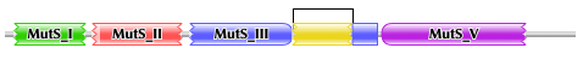
Pfam and SMART were used to characterize and image the protein domains of the MSH2 protein. SMART confidently identified two protein domains in the MSH2 protein, the MUTSd and MUTSac domains (Figure 1). The confidence of these protein domains is shown in Figure 2. Pfam identified five protein domains in the MSH2 proteins, MutS I - V (Figure 3).
Pfam and SMART did not describe in detail the function of each of the protein domains, so I used PubMed to find studies that characterized these functions. Protein domain functions are summarized in Figure 4. Briefly, MutS I is a mismatch binding domain that recognizes and binds DNA mismatches in newly replicated DNA. MutS II, III, and IV are connector, lever, and clamp domains, respectively, that play different roles in holding the DNA that is to be repaired. MutS V is an ATPase domain. Within these protein domains, there are also necessary interaction regions. The interaction regions that binds MSH3 and MSH6 to form MutS-alpha and MutS-beta are located within protein domains MutS III, IV, and V. The interaction region that binds the exonuclease EXO1 is mostly located within the MutS V protein domain.
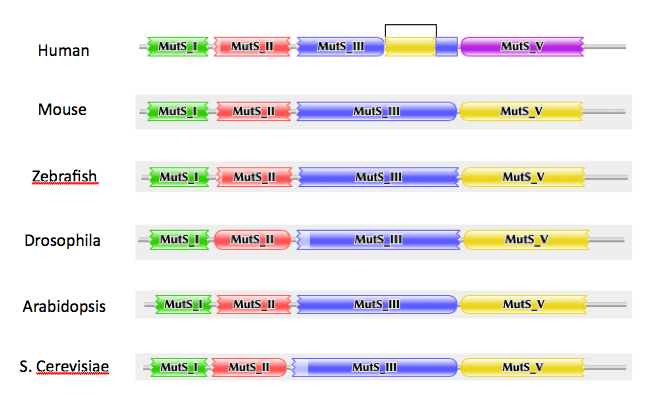
Conservation of protein domains in the human MSH2 protein was analyzed. Using Pfam, the protein domains of model organism homologs of the human MSH2 protein were identified (Figure 5).
Analysis
Protein domains of the human MSH2 protein were identified using Pfam and SMART, and each of their functions were characterized using a PubMed literature search. MSH2 model organism homolog analysis shows that the the protein domains of the MSH2 protein are very well conserved. This indicates that these proteins should behave very similarly in vivo, and thus any of these model organisms could be used to study the MSH2 protein.
References
1. Anne Lützen, Niels de Wind, Dubravka Georgijevic, Finn Cilius Nielsen, Lene Juel Rasmussen. Functional analysis of HNPCC-related missense mutations in MSH2. Mutation Research. 2008. 645;44–55.