This page was produced as an assignment for Genetics 677, an undergraduate course at UW-Madison.
Geographical MSH2 Mutations
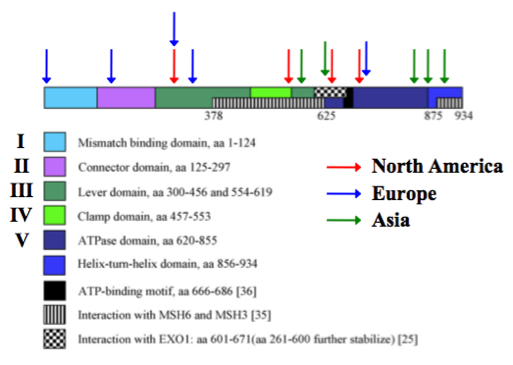
Characterization of the wild type MSH2 gene and protein is important for understanding the role of MSH2 is HNPCC, but it is also important to study mutated forms of MSH2 as well. For this reason, I first wanted to determine where in the MSH2 gene mutations commonly occur that give rise to HNPCC. In order to do this, I conducted a mini meta-analysis using PubMed and Web of Knowledge [1-13]. Any mutation that decreases the function of the MSH2 protein would result in a higher mutation rate and thus a higher likelihood of mutations occurring in the necessary genes to cause colon cancer. It turns out that mutations can occur in all domains of the MSH2 protein. In HNPCC patients, nonsense mutations seem to be most common since they are the most likely to cause loss of MSH2 function. Deleterious missense mutations can also occur causing HNPCC. Since so many different mutations have been found in the MSH2 gene, I decided to focus only on the most common missense mutations, that is missense mutations that were found independently in two or more studies or ones that were found in multiple cases within the same study. I also limited my analysis to missense that are likely to be deleterious, namely, mutations that changed amino acid side-chain polarity or charge. Mutations that meet these requirements are mapped in Figure 1.The studies included in the meta-analysis organized HNPCC cases by geographical region, so I did the same thing.
As shown in Figure 1, the preferential location of potential deleterious MSH2 mutations in HNPCC patients seems to be influenced by the geographical region in which the patient resides. Europeans seem to be more prone to mutations near the N-terminal end of MSH2, Asians seem to be more prone to mutations near the C-terminal end of MSH2, and North Americans seem to be more prone to mutation in the middle of the MSH2 gene. From these initial findings, I want to ask the testable research question stated below.
Question: Does geographic region correlate with severity of HNPCC?
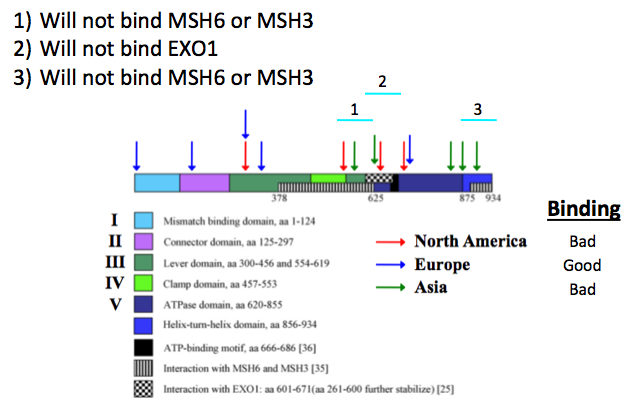
Hypothesis: Based my findings shown in Figure 1, I hypothesize that HNPCC patients in Europe will have the least severe form of HNPCC because the most common MSH2 mutations that are found in these patients are not located in essential MSH3, MSH6, or EXO1 interaction domains. HNPCC patients in Asia and North America, however, tend to have MSH2 mutations in MSH3, MSH6, and EXO1 interaction domains that may inhibit or prevent these interactions and cause loss of MSH2 function.
In order to test this hypothesis, the individual function of these mutant MSH2 proteins must be test. To reiterate, a less functioning MSH2 would likely result in a more severe HNPCC predisposition to cancer. Function of mutant MSH2 proteins will be tested by:
1) MSH2 binding affinity with protein partners
2) Mutator Phenotype – High mutation rate
Hypothesis: Based my findings shown in Figure 1, I hypothesize that HNPCC patients in Europe will have the least severe form of HNPCC because the most common MSH2 mutations that are found in these patients are not located in essential MSH3, MSH6, or EXO1 interaction domains. HNPCC patients in Asia and North America, however, tend to have MSH2 mutations in MSH3, MSH6, and EXO1 interaction domains that may inhibit or prevent these interactions and cause loss of MSH2 function.
In order to test this hypothesis, the individual function of these mutant MSH2 proteins must be test. To reiterate, a less functioning MSH2 would likely result in a more severe HNPCC predisposition to cancer. Function of mutant MSH2 proteins will be tested by:
1) MSH2 binding affinity with protein partners
2) Mutator Phenotype – High mutation rate
Experiment #1: Does missense mutant MSH2 bind to MSH6, MSH3, and EXO1?
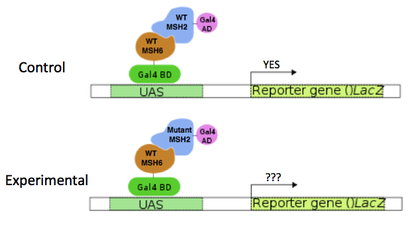
Figure 2. Diagram of the proposed yeast 2-Hybrid experiment.
I first propose to use a yeast 2-Hybrid experiment to test whether mutant MSH2 proteins can interact with MSH3, MSH6, and EXO1. An animation describing the yeast 2-Hybrid method can be found HERE. Briefly, physical interaction of the two proteins of interest is required and causes the transcription of a reporter gene (Figure 2). Each mutant MSH2 protein will be tested using this method for binding with the interacting proteins MSH3, MSH6, and EXO1. These physical interactions are necessary for proper MSH2 function. Wild type MSH2 will be used as a control.
Expected Results
My expected results are shown in Figure 3. I expect that, on average, European mutant MSH2 proteins would interact with MSH3, MSH6, and EXO1 as well as wild type MSH2. On the other hand, I expect that Asian and North American mutant MSH2 proteins would interact poorly with MSH3, MSH6, and EXO1 compared to wild type MSH2 because mutations in these proteins occur in the necessary interacting domains.
My expected results are shown in Figure 3. I expect that, on average, European mutant MSH2 proteins would interact with MSH3, MSH6, and EXO1 as well as wild type MSH2. On the other hand, I expect that Asian and North American mutant MSH2 proteins would interact poorly with MSH3, MSH6, and EXO1 compared to wild type MSH2 because mutations in these proteins occur in the necessary interacting domains.
Experiment #2: Does missense mutant MSH2 higher mutation rate (mutator phenotype)?
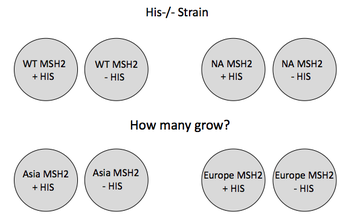
Figure 4. Diagram of the proposed mutator experiment
A histidine null (His-/-) S. cerevisiae strain will be utilized in this experiment. Many His-/- strains will be mutated to have corresponding MSH2 mutations as the HNPCC patients. These strains will then be plated on media with and without histidine. Colonies that grow without histidine have acquired a reversion mutation. Reversion rate of mutant MSH2 strains with a His-/- background to a His(+) phenotype is correlated with the functional efficiency of the MSH2 protein. A higher reversion rate means a higher mutation rate which means a less functional MSH2 protein and a more severe HNPCC. Strains with wild type MSH2 protein will be used as a control. This experiment is summarized in Figure 4.
Expected Results
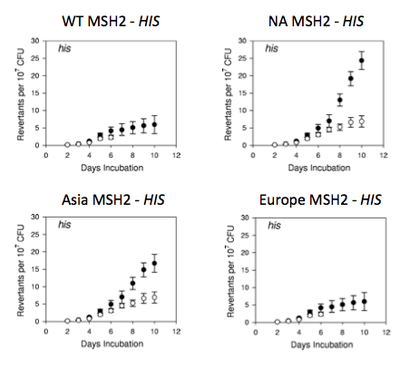
Figure 5. Expected results of the proposed mutator experiment.
I would expect the strain with wild type MSH2 to have a relatively low reversion rate since its DNA mismatch repair machinery should be fully functional. On average, I would expect strains with European MSH2 mutations to be a little less functional than the wild type strain because the mutations affect DNA binding domains but not essential MSH3, MSH6, EXO1, or ATP binding domains. I would also expect strains with Asian or North American MSH2 mutations to be much less functional than the wild type strain, and perhaps nonfunctional, because these mutations affect necessary MSH3, MSH6, EXO1, and ATP binding domains. These results are shown in Figure 5.
Analysis
The proposed research studies should reveal if there is a correlation between geographic region and severity of HNPCC. It is important to note, however, that the initial mutation finding are likely biased. They are not only biased in which published results I found and included in my meta-analysis, but they are also biased in which countries have the resources and will to study HNPCC and in which patients were analyzed. This is especially true in the case of HNPCC since there are so many different MSH2 mutations that can cause HNPCC. Thus, the subset of mutations that are mentioned and analyzed here may not accurately reflect the global population.
The identification of a correlation between geographic region and severity of HNPCC would be important for physicians in order to make better treatment decisions. It is also noteworthy to mention that this study cannot distinguish between environmental factors in the geographic region or genetic factors of that particular ethnic background that cause these different MSH2 mutations. It is likely to be the latter case, but different analyses would be required to test this.
The proposed research studies should reveal if there is a correlation between geographic region and severity of HNPCC. It is important to note, however, that the initial mutation finding are likely biased. They are not only biased in which published results I found and included in my meta-analysis, but they are also biased in which countries have the resources and will to study HNPCC and in which patients were analyzed. This is especially true in the case of HNPCC since there are so many different MSH2 mutations that can cause HNPCC. Thus, the subset of mutations that are mentioned and analyzed here may not accurately reflect the global population.
The identification of a correlation between geographic region and severity of HNPCC would be important for physicians in order to make better treatment decisions. It is also noteworthy to mention that this study cannot distinguish between environmental factors in the geographic region or genetic factors of that particular ethnic background that cause these different MSH2 mutations. It is likely to be the latter case, but different analyses would be required to test this.
References
1. Lee S-C, Guo J-Y, Lim R, Soo R, Koay E, Salto-Tellez M, Leong A, Goh B-C. Clinical and molecular characteristics of hereditary non-polyposis colorectal cancer families in Southeast Asia. Clin Genet. 2005. 68;137–145.
2. Wei W, Liu L, Chen J, Jin K, Jiang F, Liu F, Ran R, Cheng Z, Shen M, Xue C, Cai S, Xu Y, Nan P. Racial Differences in MLH1 and MSH2 Mutation: An analysis of yellow race and white race based on the insight database. Journal of Bioinformatics and Computational Biology. 2010. 8(1);111-125.
3. Anja Wagner, Alicia Barrows, Juul Th. Wijnen, et al. Molecular Analysis of Hereditary Nonpolyposis Colorectal Cancer in the United States: High Mutation Detection Rate among Clinically Selected Families and Characterization of an American Founder Genomic Deletion of the MSH2 Gene. 2003. Am. J. Hum. Genet. 72:1088–1100.
4. Young Mee Kim, Chang Gyu Choe, Somi Kim Cho, In Ho Jung, Won Young Chang, Moonjae Cho. Three novel germline mutations in MLH1 and MSH2 in families with Lynch syndrome living on Jeju island, Korea. BMB reports. 2010. 43(10);693-697.
5. Elisabeth Mangold, Constanze Pagenstecher, Waltraut Friedl, et al. Spectrum and frequencies of mutations in MSH2 and MLH1 identified in 1,721 german families suspected of hereditary nonpolyposis colorectal cancer. Int. J. Cancer. 2005. 116;692–702.
6. Kurzawski G, Suchy J, Lener M, Kłujszo-Grabowska E, et al. Germline MSH2 and MLH1 mutational spectrum including large rearrangements in HNPCC families from Poland (update study). Clin Genet. 2006. 69;40–47.
7. Yun-HeeChoi, MichelleCotterchio, GailMcKeown-Eyssen, et al. Penetrance of colorectal cancer among MLH1/MSH2 carriers participating in the colorectal cancer familial registry in Ontario. Hereditary Cancer in Clinical Practice 2009, 7:14.
8. George Chong, Jonathan Jarry, Victoria Marcus, et al. High Frequency of Exon Deletions and Putative Founder Effects in French Canadian Lynch Syndrome Families. HUMAN MUTATION Mutation in Brief. 2009. 30:E797-E812.
9. Lise Lotte Christensen, Bo E Madsen, Friedrik P Wikman, et al. The association between genetic variants in hMLH1 and hMSH2 and the development of sporadic colorectal cancer in the Danish population. BMC Medical Genetics 2008, 9:52.
10. Francesco Baudia, Giuseppina Fersinia, Annamaria Lavecchiaba, et al. A novel missense germline mutation in exon 2 of the hMSH2 gene in a HNPCC family from Southern Italy. Cancer Letters. 2005. 223;285–291.
11. Francesca Bianchi, Eva Galizia, Emilio Porfiri, et al. A missense germline mutation in exon 7 of the MSH2 gene in a HNPCC family from center-Italy. Familial Cancer. 2007. 6:97–102.
12. Tang R, Hsiung C, Wang J-Y, et al. Germ line MLH1 and MSH2 mutations in Taiwanese Lynch syndrome families: characterization of a founder genomic mutation in the MLH1 gene. Clin Genet. 2009. 75;334–345.
13. Wenqian Wei, Fangqi Liu, Lei Liu, et al. Distinct mutations in MLH1 and MSH2 genes in Hereditary Non-polyposis Colorectal Cancer (HNPCC) families from
China. BMB reports. 2011. 44(5);317-322.
14. Anne Lützen, Niels de Wind, Dubravka Georgijevic, Finn Cilius Nielsen, Lene Juel Rasmussen. Functional analysis of HNPCC-related missense mutations in MSH2. Mutation Research. 2008. 645;44–55.